GC IMAGE - GC Image LCxLC Edition - User Guide 2025R2
New Features and Improvements in Version 2025
GC Image LCxLC Edition 2025 has many new features and improvements. See
the GC Image LCxLC Edition User Guide for full
documentation.
What's New in Version 2025 Release 2 (November 2025)
Version 2025R2 includes new and improved I/O support for LECO, Bruker, and JEOL data formats:
- Expanded LECO ChromaTOF .PEG and .SMP Support:
- The software now supports reading modulation information (single and variable modulation) when importing .PEG files.
- The software now supports importing additional versions of .PEG files, including those acquired with dual FID/SCD detectors.
- The software now supports importing .SMP files generated from .PEG source files.
- The software now supports importing certain newer versions of .SMP files that contain non-spectral signals.
- The software now supports importing Bruker timsTOF .TSF data files.
- The software now includes JEOL's msAxelDataReader version 1.4, adding support for data files acquired with JEOL AccuTOF GC-Alpha (JMS-T2000GC).
Version 2025R2 includes a newly designed UI for the operation
journal, which makes it easier to verify audit trails and
quickly find previously used processing settings.
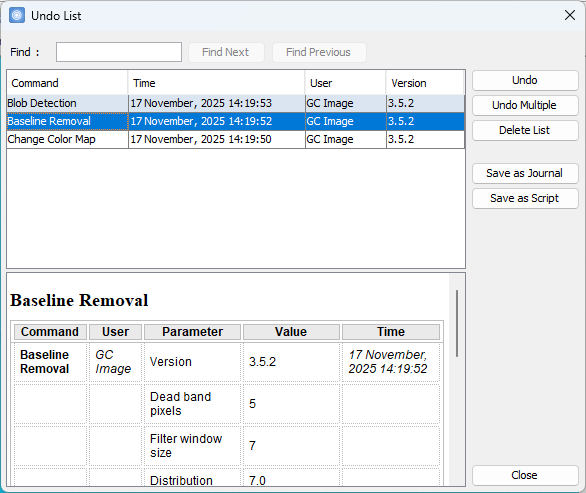
- Improved List Operations and Journal UI
Edit > List
Operations and View
> Journal now present a redesigned dialog with:
- A table layout for clearer visibility of
operations,
- A Find bar for quick searching,
- A detail pane that displays parameter details
of the selected operation.
- Undo Multiple Operations at Once
A new Undo
Multiple button has been added to Edit > List
Operations. Users can now select and revert a sequence
of operations up to and including the chosen row.
Version 2025R2 includes an updated
Method Editor and expanded set of operations that
provide a more complete and customizable workflow. These
improvements enhance usability, streamline template handling,
and simplify saving key analysis tables.
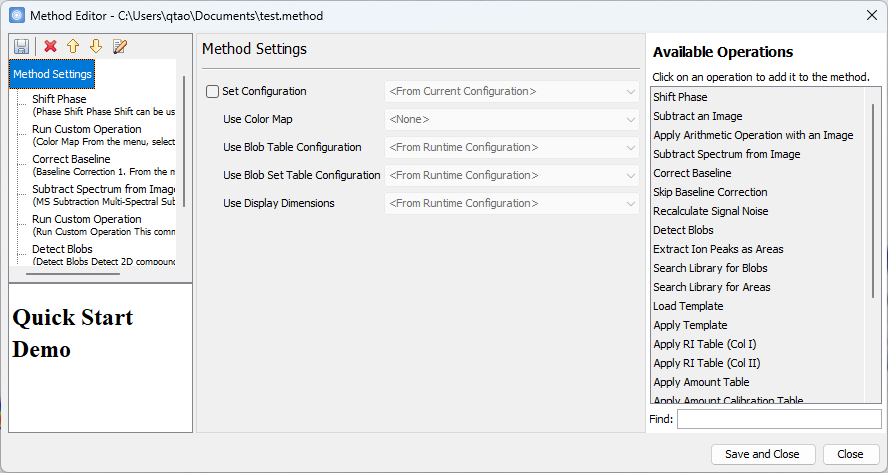
- The Method Editor now
has a more polished interface, including a new Find bar
that enables searching and filtering across available
operations.
- Template method operations
can now be more flexible and allow users to combine
multiple templates and apply templates after other alignment
or match operations.
- Apply Template operation includes two new options
From Runtime Image (to use the template
already in the image) and None for Match Type (to
apply the template as-is without matching).
- Load Template operation allows specifying a
template at runtime or copying from the
currently open image, with a Load Type parameter to
overwrite or append to the existing template.
- New operations allow saving the Blob Table, Area Table, and Blob
Set Table to Excel or CSV files. This makes it easier to
export compound analysis results for reporting or further processing.
- New Run Custom Operation
steps can be incorporated directly into a method to invoke
executable plugin commands, enabling integration of external
tools and extending method workflows with custom
functionality.
 |
Version 2025R2 now supports interactive methods that enable
standardizing workflows that require human input, with
step-by-step execution and custom instructions at each stage.
This approach helps retain and share expert know-how, while
providing greater control and flexibility during method runs.
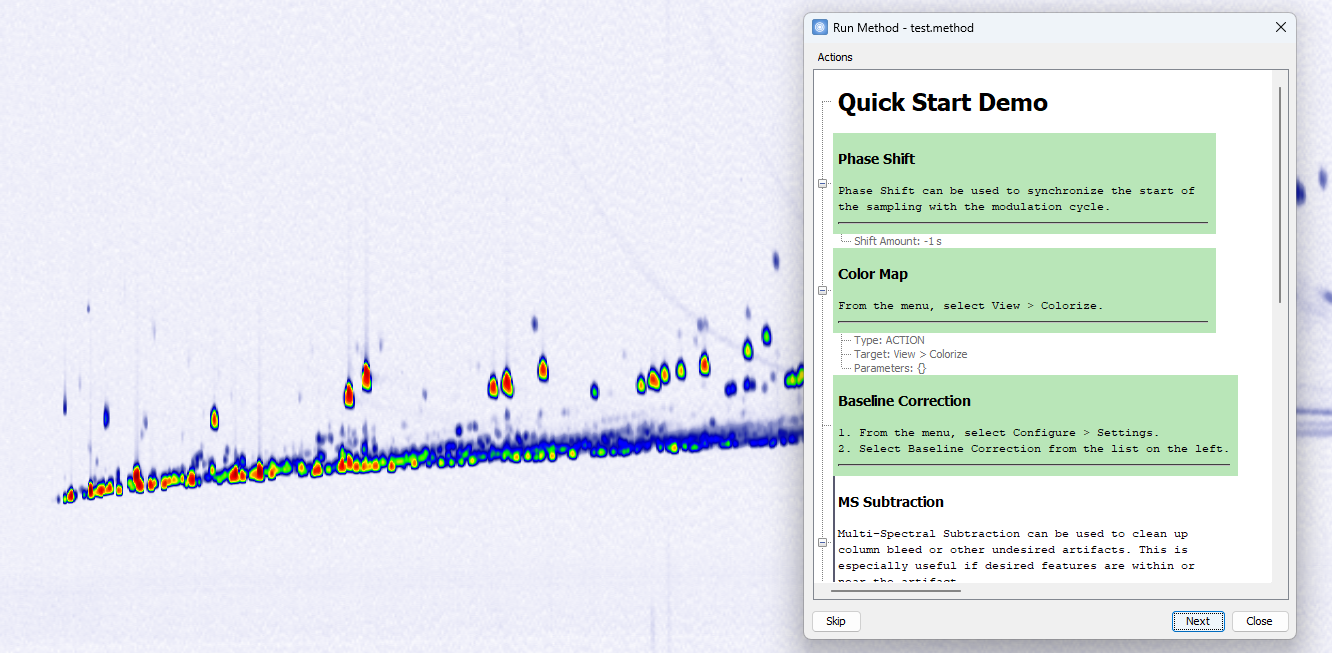
- Support for customizable titles and descriptions in method steps
The Method Editor now
supports adding descriptive text to the entire method and to
individual steps. These descriptions accept basic HTML
formatting, enabling headings, styled text, and lists, and
are displayed before and during interactive runs.
- Support for step-by-step interactive execution
Methods can
now be executed in Interactive Mode
as an alternative to running them all at once. This mode
provides greater control by allowing execution step-by-step,
with opportunities for manual verification and adjustments
between steps.
- Support for pause and menu actions via Run Custom Operation
The new Run Custom Operation
steps can be incorporated directly into a method to pause
execution or invoke any menu command by specifying its path,
enabling more flexible and guided workflows.
 |
Version 2025R2 now includes an enhanced adduct library that allows
users to import their own adduct definitions, making it easier
to tailor analyses to specific experimental needs.
- Support for importing user-defined adducts
The new
Configure > Adducts Library
provides a centralized interface for managing the complete
list of available adducts, with options to view, search,
import, and restore entries. It supports importing custom
adducts from CSV or Excel files, enabling laboratories to
extend the library with specialized adducts.
- Updated default adduct list
The default adduct list has
been expanded to include additional positive ion species
(M+, M+2, M+3, M+H-2H2O, M+H-H2O) and negative ion species
(M-, M-2, M-3, M+HCOO, M+CH3COO).
Version 2025R2 introduces several enhancements across reporting
and tool management, improving flexibility, compatibility, and
user experience.
- Tools > Slice Report for Boiling Range Distribution
now supports the use of an external calibration table stored
in a CSV or Excel file.
- Both Summary Report
and Filter Report
can now be saved directly as HTML files, with HTML set as
the default file type.
- The Save Summary Report method operation now includes a
Preferred Report File Type parameter, allowing selection of
HTML or Excel in addition to XML file type.
- The Plugin
Manager now has an improved interface with
a redesigned plugin table that highlights key columns, a
Filter bar for quick searching by name, and an
enhanced import process that prompts whether to remove an
existing plugin before updating.
What's New in Version 2025 Release 1 (June 2025)
Version 2025R1 includes a significant update to the ion peak
detection algorithm with improving reliability and precision. This
updated algorithm enhances ion peak merging and noisy peak filtering,
delivering more accurate peak detection and deconvolved spectra. It
replaces the previous version and is now the default.
In Version 2025R1, calculations using CLIC functions, FEATUREATBLOB()
and IDOF(), especially when referencing a specific identified
blob like an ISTD, now run faster than ever. This enhanced speed
allows for more complex and flexible normalization and calculations,
for example, Volume/FEATUREATBLOB(IDOF("Biphenyl-d10", "Blob"),
"Volume"), to be executed smoothly in Blob Table.
Version 2025R1 introduces a smarter alignment marker selection process in the
Compare Images Tool, enhancing accuracy and
reliability. The updated algorithm refines initial matching by pairing
identically named blobs across images while prioritizing consistency
with the predicted transform. This ensures the most stable and
globally reliable transformation.
Additionally, a new Only Use Included option allows users to
exclusively utilize included blobs when initializing alignment
markers, providing greater control over the selection process.
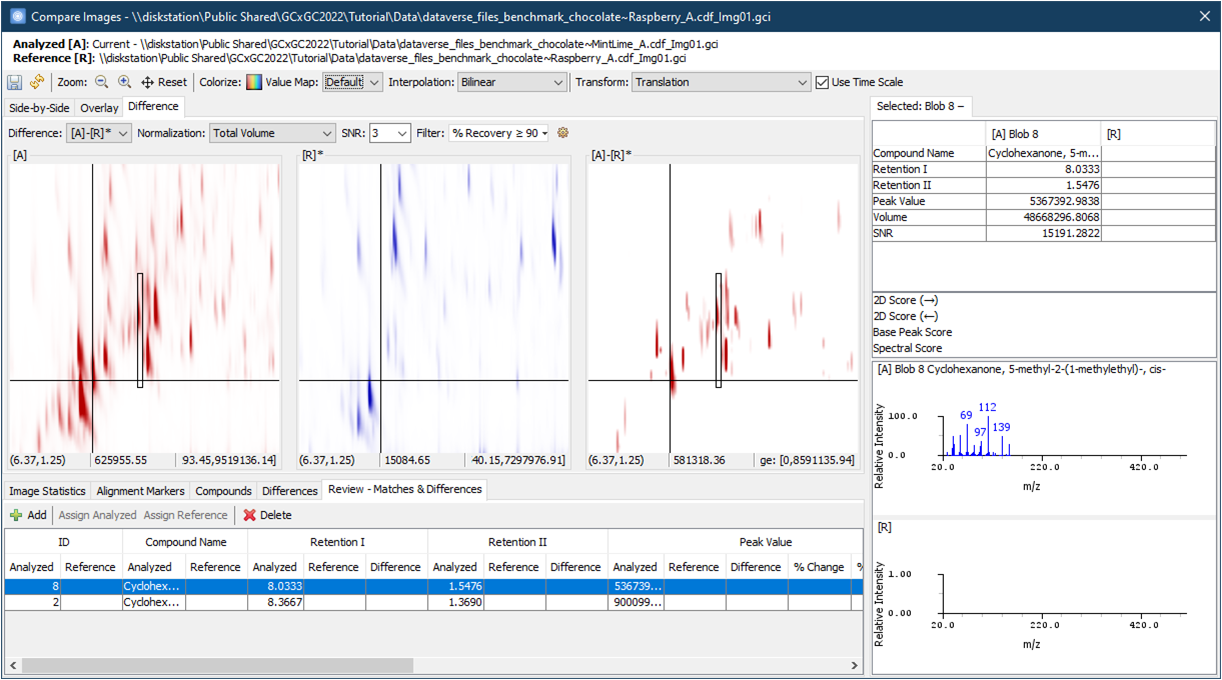
Version 2025R1 introduces Difference View, a powerful tool
designed to quickly identify and analyze differences between two
chromatograms. This directional comparison computes a pixel-by-pixel
difference, highlighting what is present in the first operand image
but absent or reduced in the second.
This new Difference View leverages precise alignment using
alignment markers and transforms, and offers advanced options
including:
- Normalization: Adjusts how the second image is normalized to
the first.
- Fuzzy Radius: Optimizes pixel matching and counters local
variations for improved accuracy.
- Relative Difference and SNR Filtering: Select None, % Recovery,
or Fold Change, to refine results while accounting for baseline
fluctuations.
In addition, a dedicated Review table displays
differentiating compounds, either matched or unmatched, that have been
reviewed and added for further analysis.
Version 2025R1 introduces new analysis tools in the
Investigator interface including:
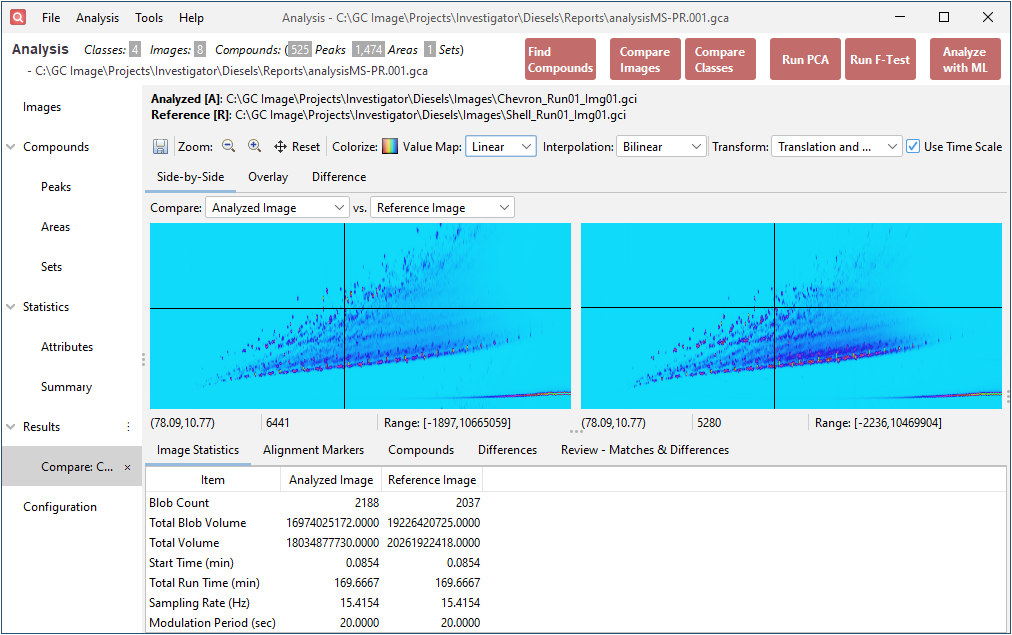
- Integrated Compare Images Support within
Analysis: Now seamlessly integrate the Compare Images
tool directly within the Analysis Actions. This
integration enables a more comprehensive comparison of two samples by
combining analytical insights with direct visual inspection of
chromatograms and peaks, ensuring thorough evaluations.
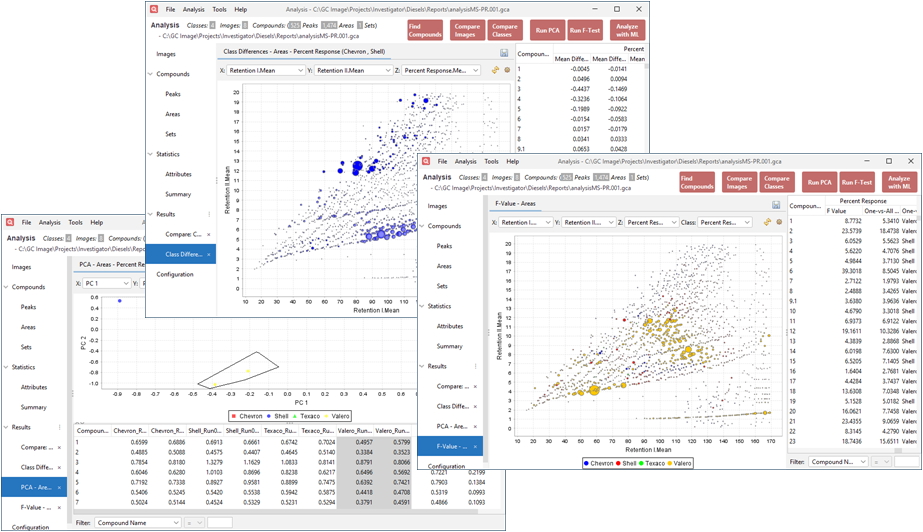
- New Pre-Configured Analysis Tools:
- Compare Classes: Generate a bubble plot of
the mean pairwise class differences for a specified attribute.
- Run PCA: Perform principal component
analysis using the selected attribute for unsupervised clustering.
- Run F-Test: Create bubble plots displaying
F-values to evaluate compound significance for distinguishing classes.
In addition, Version 2025R1 introduces a refreshed interface for
Investigator to optimize usability and visualization:
- Better Chromatogram View displays a sharper
and more visually appealing chromatogram snapshot at any viewing
size.
- Redesigned Analysis Navigation uses the new
side-bar system, replacing the old tab-based interface for a more
organized and intuitive experience.
Contents
Next: Introduction
GC Image™ LCxLC Edition User Guide ©
2001–2025 by GC Image, LLC, and the University of Nebraska.